Revised Protocol of Non-Invasive Prenatal Test More Accurately Detects Angelman, Other Genetic Diseases, Study Shows
Written by |
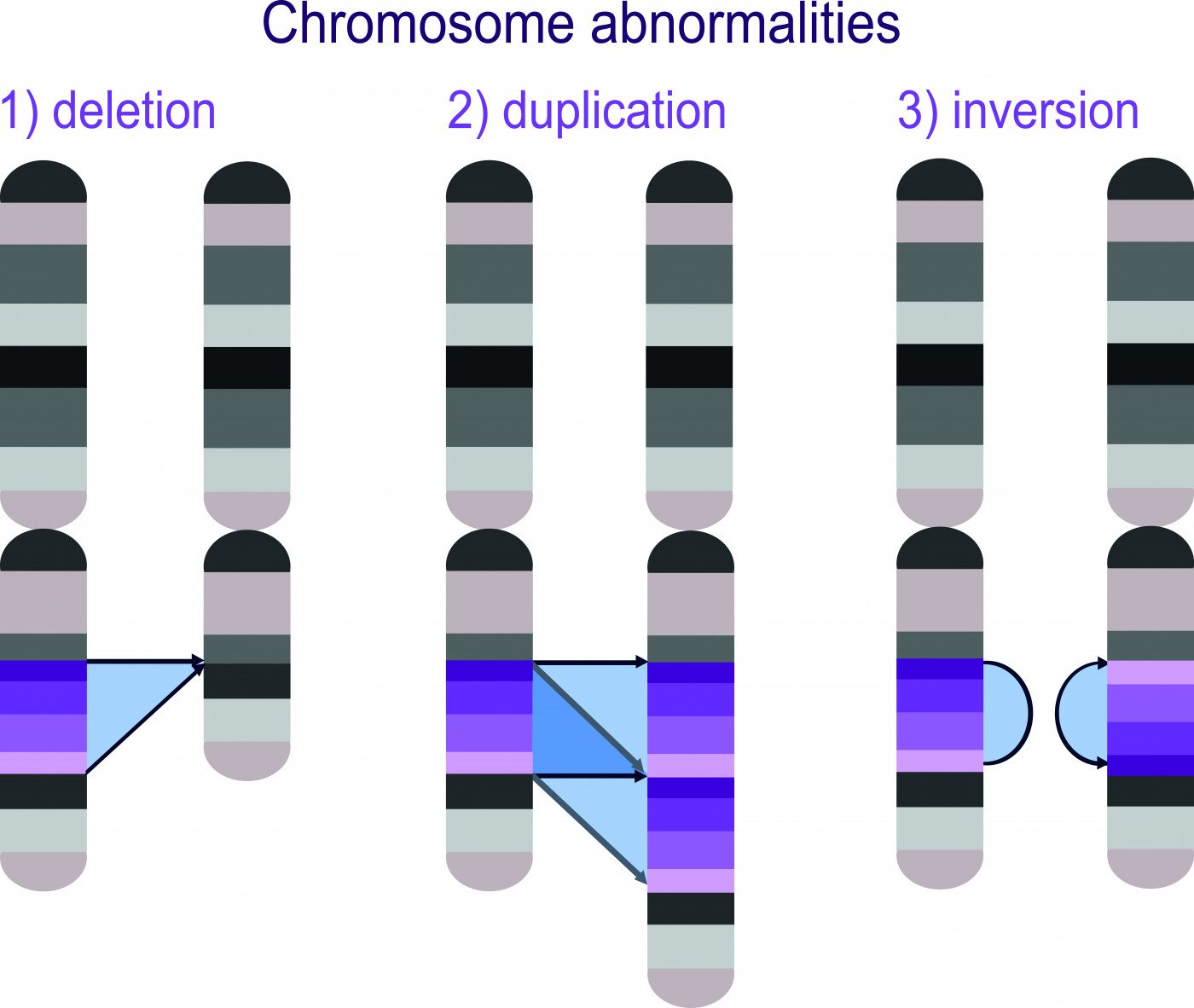
The genomic deletions associated with Angelman syndrome, 1p36, cri-du-chat, and Prader-Willi can be more accurately detected with the revised protocol of a non-invasive prenatal test, a new study shows.
The test analyzes tiny pieces of the fetus’ DNA, which circulate freely in the mother’s blood, and detects small changes in nucleotides (the building block of DNA) that might flag for a certain genetic disease.
The study, titled “Clinical Experience with a Single-Nucleotide Polymorphism-Based Noninvasive Prenatal Test for Five Clinically Significant Microdeletions,” was published in Clinical Genetics.
Chromosomal abnormalities in fetuses can encompass both whole chromosome abnormalities, such as in Down’s syndrome, and microscopic abnormalities, such as in Angelman’s syndrome. Microscopic abnormalities tend to be copy-number variants (CNVs), which may either be microdeletions in the genetic code or microduplications.
The prevalence of these microdeletions can lead to clinically significant phenotypes (the observable characteristics of an organism). When a structural abnormality indicating a microdeletion is observed on an ultrasound, the mother typically is referred to undergo a chromosomal analysis (CMA). However, this invasive procedure involves obtaining fetal cells for analysis, and comes with a risk of fetal loss. Therefore, many expecting mothers decline CMA.
One non-invasive procedure, called SNP-based non-invasive prenatal testing (NIPT), involves analyzing cell-free DNA that is present in the maternal plasma to help identify microdeletions. This technique can screen for five different microscopic chromosomal abnormalities including Angelman (maternal 15q11–q13), 22q11.2, 1p36, cri-du-chat (5p15.3), and Prader-Willi (paternal 15q11–q13) syndromes.
But one of the issues with this test is the high level of false positive rates. Studies have shown that re-sequencing at a higher depth of read (HDOR) can improve the false-positive rate.
That led researchers at the University of Connecticut to examine the performance of the HDOR modification to the SNP-NIPT test, as well as a better quality threshold to determine the prevalence of these five microdeletions in the population.
They conducted a one-year retrospective analysis of the test performance for 80,449 women who were referred for microdeletion screening. Among them, 42,326 were screened for all five microdeletions and the rest were screened only for 22q11.2, which is the most common microdeletion.
Results showed that the positive predictive value for the original test — the SNP-NIPT alone — was 15.7% for 22q11.2 deletion syndrome, and 5.2% for the other four disorders combined. Positive predictive value is the probability that patients with a positive screening test actually have the disease. When re-conducting the test with the revised protocol including the HDOR, these positive predictive values increased to 44.2% for 22q11.2 and 31.7% for the other four microdeletions.
Furthermore, initially the false positive rate for the SNP-NIPT test was 0.33% for 22q11.2 deletion syndrome. Upon re-conducting analysis with the revised protocol, the false-positive rate decreased to 0.07%. Additionally, the false positive rate for the other four disorders combined decreased from 0.56% to 0.07%.
Researchers determined that the minimal prevalence of these five diseases in the population were 1/1,255 for 22q11.2 deletion syndrome and 1/1,464 for 1p36, cri-du-chat, Prader-Willi, and Angelman syndromes combined in the referral population. The combined minimum prevalence of all five diseases in the referral population was 1/676.
“The performance of the original screening protocol for 22q11.2 deletion syndrome in this cohort is similar to that demonstrated previously. We also show that applying a stricter quality-control confidence threshold and reflexively sequencing high-risk samples to a higher depth of read with the revised screening protocol reduced FPRs (false positive rates), and increased PPVs (positive predictive value) substantially. As a result of these performance improvements, the revised protocol has been implemented commercially,” the authors concluded.

